Ellf: Your virtual NLP engineer
Ellf (short for Explosion Large Language Framework) is an interactive, AI-powered assistant for Natural Language Processing (NLP) and machine learning projects. It helps you design solutions, break down problems, iteratively develop data and train, evaluate and deploy your own models. With Ellf, your team can take back control and build custom and fully private AI solutions in-house.
Ellf integrates with your coding assistant like Claude Code and makes it proficient at planning and developing NLP solutions. It also includes a powerful and fully data-private cluster that runs locally or in the cloud and makes it easy to execute annotation tasks, auto-annotation agents, training experiments and more, and collaborate on development with your team.
Ellf is currently in private beta
Features and APIs may change as we iterate based on feedback. To join the waitlist and be notified once Ellf is available, you can sign up here. We’re also currently looking for beta partners who need help with their NLP projects, especially around tasks like information extraction. If that’s you, sign up for the waitlist and include details about your project, or get in touch via email.
First steps
Install the Ellf client
Download the CLI and connect Ellf to your coding assistant like Claude Code.
Set up your cluster
Launch your private data processing cluster locally or in the cloud.
Explore the platform
Get started and set up your first projects, tasks, actions and agents.
Once you’ve installed Ellf and connected it to your coding assistant, you can start by describing your project and asking it questions on strategy and next steps. If you already know what you want to do, you can also call the individual modules directly – for example, the /ellf-project command starts the project planning module and /ellf-annotate helps you design data annotation tasks.
› I need to detect mentions of adverse drug reactions in medical literature abstracts. My team has
about 500 papers but nothing annotated yet. What’s the best approach?
⏺︎Let me analyze your requirements and put together a project plan for you.To keep track of your project, Ellf will create detailed project plans. These are Markdown documents that are continuously updated and outline the end-to-end development plan, including step-by-step workflows and higher-level strategic goals.
NLP Project Plan: ADR Detection in Medical Literature
Problem Statement
Detect and extract structured adverse drug reaction (ADR) mentions from ~500 biomedical paper abstracts (currently in PDF/mixed format), producing typed drug-ADR pairs suitable for ingestion into a pharmacovigilance database.
Pipeline Overview
PDF/mixed source
└─> [Phase 0] Text extraction (Docling / pdfplumber + spacy-layout)
└─> [1] DRUG NER (scispaCy BC5CDR baseline → fine-tune if needed)
└─> [2] ADVERSE_REACTION NER (same model, separate eval pass)
└─> [3] Relation linking (sentence co-occurrence + negation → NLI reframe if needed)
└─> [4] Normalization (DrugBank/RxNorm, MedDRA) — business logic, not ML
Data Strategy
- ~500 abstracts in PDF/mixed formats — text extraction required before NLP work
- Nothing annotated yet
- Plan: LLM pre-annotation (GPT-4 via spacy-llm) → expert correction in Prodigy
- Set aside ~100 abstracts as held-out eval set before any annotation begins
- Target: ~300–400 annotated abstracts for NER training
Annotation Plan
Schema
DRUG— drug/substance mention spans (generic, not role-specific)ADVERSE_REACTION— adverse reaction term spans (generic)- Relations: drug-reaction pairs (separate pass after NER is stable)
Schema principles
- Do NOT use composite labels like "DRUG_CAUSING_ADR" — couple NER with role assignment is a mistake
- Normalization to DrugBank / MedDRA is business logic downstream, not part of the annotation schema
Workflow
- Pilot: 50–100 abstracts, domain expert annotates DRUG spans only (correction of BC5CDR predictions)
- Evaluate BC5CDR baseline on same set — decide how much fine-tuning is needed
- ADVERSE_REACTION pass (single-label correction mode)
- Relation pass: link drug-reaction pairs within sentences
- Train first model after pilot, run training curve, validate learnability before scaling
Team
- 1–2 domain experts (clinicians / pharma researchers) — ideal small expert team
- Prodigy on Ellf cluster for annotation
Evaluation Strategy
- Hold out ~100 abstracts as a fixed test set before training starts — never train on these
- Evaluate NER with token-level and span-level F1, split by entity type
- Rule-of-10: distinguish 85% vs 90% accuracy needs ~1000 span-level eval examples — plan accordingly
- Relation evaluation: precision/recall on drug-ADR pairs (exact match: both spans + link correct)
- Run training curves after pilot to decide whether more annotation helps or schema needs revision
- Baseline: evaluate scispaCy BC5CDR out-of-the-box before annotating anything
Roadmap
Phase 0 — Text extraction (prerequisite)
- Extract clean abstract text from PDFs
- Tools: Docling, pdfplumber, or spacy-layout
- Output: plain text files per abstract, ready for annotation
Phase 1 — Baseline evaluation
- Run scispaCy en_ner_bc5cdr_md on 50 sample abstracts
- Measure what it gets right on DRUG and ADVERSE_REACTION before committing annotation budget
Phase 2 — NER annotation + model
- LLM pre-annotation → expert correction in Prodigy (ner.correct)
- Single-label passes: DRUG first, ADVERSE_REACTION second
- Train after pilot, run training curve, scale only if curve is still rising
Phase 3 — Relation extraction
- Start with rule-based co-occurrence + negation detection as baseline
- Escalate to NLI reframe ("drug X caused reaction Y" → entailed/neutral/contradicted) if needed
Phase 4 — Normalization (business logic)
- Map DRUG spans to DrugBank / RxNorm
- Map ADVERSE_REACTION spans to MedDRA
- This is a lookup layer, not a model — keep it separate from NLP components
Next Steps
- Solve text extraction (Phase 0) — this is a blocker for everything else
- Evaluate scispaCy BC5CDR on a sample before annotating
- Set up Prodigy annotation task on Ellf with LLM-assisted pre-annotation
NLP development typically involves running many different things: data processing, annotation, training experiments, evaluation and more, often iteratively and in parallel. To make this easier, Ellf can automatically start and monitor these processes and run them in a data-private cluster, either locally or on your own cloud account. This means that no data ever has to leave your servers.
The platform lets you view your tasks, actions, agents and services, as well as data assets on your cluster, and collaborate on them with your team. You can interact with Ellf and manage resources via the ellf command-line interface or visually in the app. The web UI also includes a command bar at the top that shows the CLI command that corresponds to the current page or view, making it easy to switch between using the app and CLI.
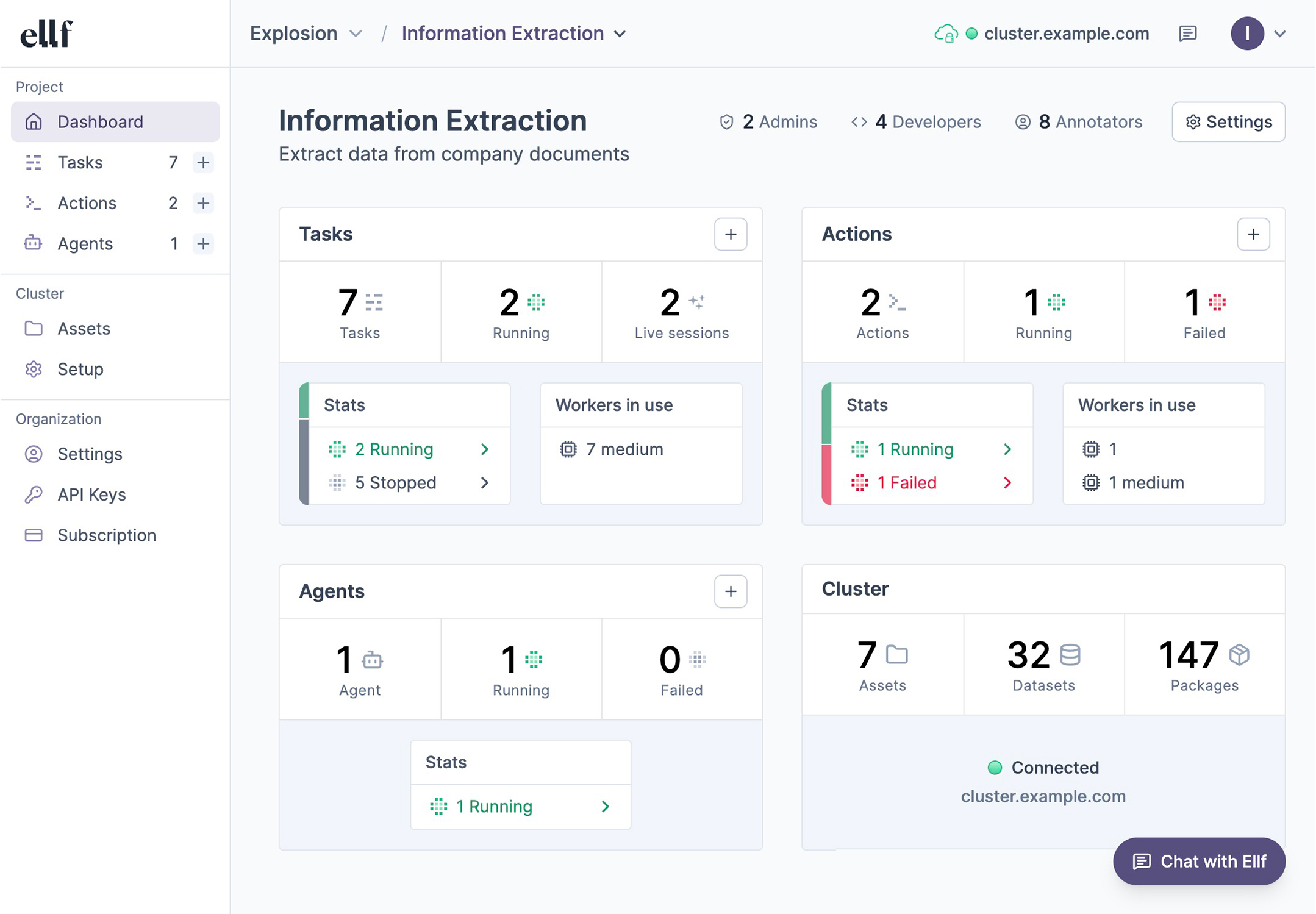
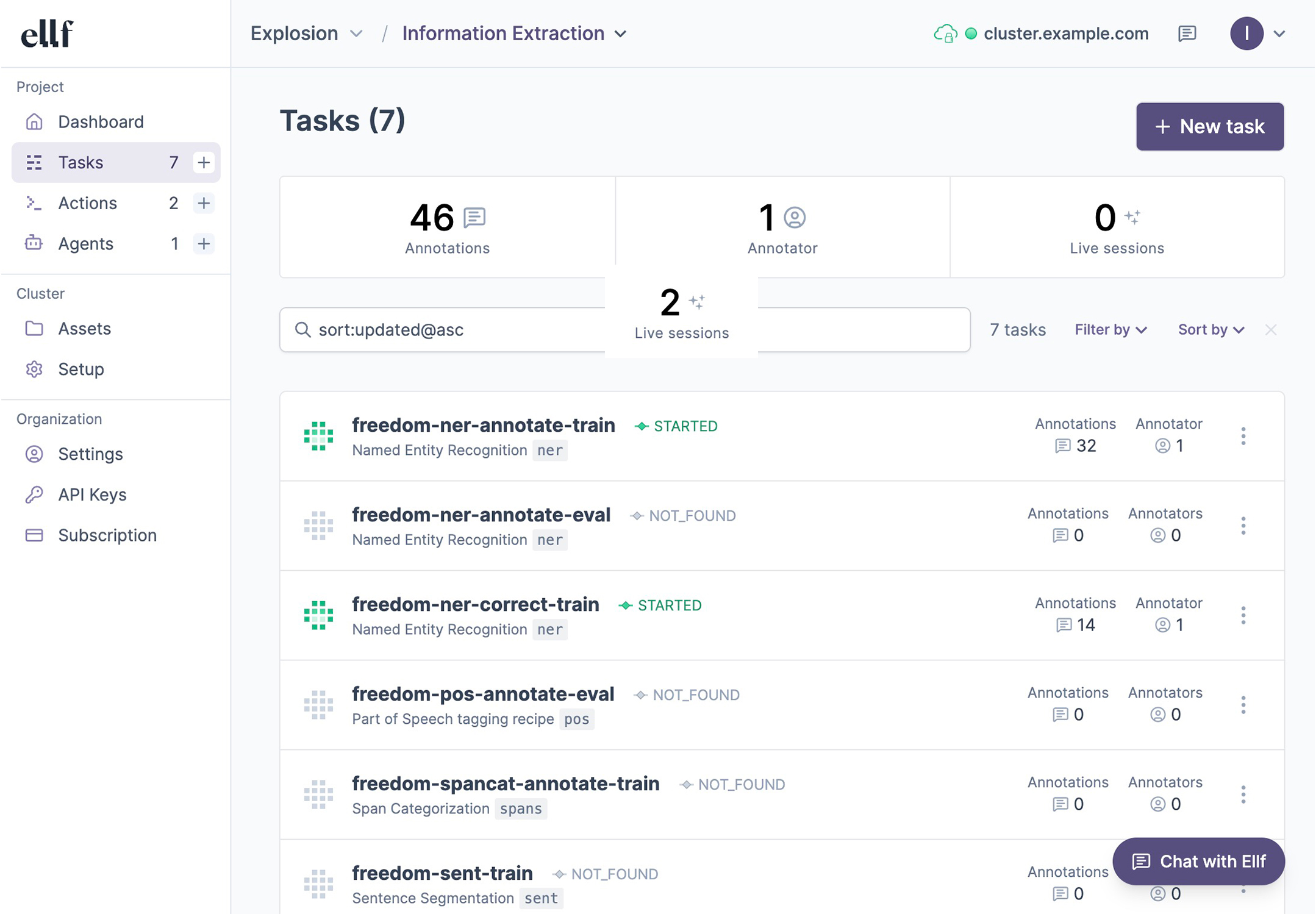
For data annotation, Ellf ships with our popular and efficient annotation tool Prodigy, a range of built-in recipes for common NLP tasks, as well as annotation agents to automate data development for you. Ellf can also help you select the right NLP components for your business problem and develop the label scheme before you scale up development.
Use Ellf to get started
If you’ve connected Ellf to your coding assistant, it will know what to do – so you can start by describing your project and requirements, and have Ellf suggest the next steps. You can also use the in-app chat to navigate the platform for you.
Architecture
To keep your data private and allow flexibly scripting the application workflows in Python, Ellf implements a novel hybrid or dual-cloud architecture, consisting of a web application and servers hosted by us, and a data processing cluster hosted by you locally or on your own cloud account and entirely under your control. The cluster stores and provides all code and data, which is executed and served directly from there.
Ellf generally stores records of data and other objects, as well as user and organization details, while your cluster stores the actual data and runs the code. Whenever data is shown or served in the web application or CLI, it comes straight from your cluster and doesn’t ever pass through servers hosted by us.
| Your cluster | Ellf |
|---|---|
| Data assets, model files, code and secrets | Records of names, IDs and paths |
| Recipe code and workers running tasks, actions, agents and services | Records of tasks, actions, agents and services, and their recipe arguments |
| Prodigy database with datasets and examples | Records of dataset names and IDs |
| Web server with Prodigy annotation app | Statistics on user accounts accessing annotation app and counts |
| References to project IDs in tasks, actions, agents and services | Metadata of created projects |
| References to user IDs in datasets | Organization and user accounts |
| In-app chat logs and project plans after migration | In-app chat logs and project plans before migration |
Human support
If you ever feel stuck or need human feedback, you can use the support module to send us your logs and have them reviewed by one of our NLP experts. Alternatively, you can also export your conversation log to a file and email it to us at support@ellf.ai.
› /ellf-support The model’s accuracy isn’t improving with more data, so I’m considering changing the label
scheme. Is this a good approach?
⏺︎Should the support request include your full session transcript, or a summary only?› Full transcript⏺︎Support request sent successfully.
- Reference ID:
a7cf3184865e4f05a8bf269ad34c767a
- Session transcript: attached
- Debug log: not found (not attached)
The Ellf team will receive your question about whether changing the label scheme is the right approach for the accuracy plateau. You’ll get a copy via email.